We check the potential funneling problem in multilevel models as described in Betancourt and Girolami (2015). Two simulation are performed.
The first example
In the first example, we have the following model: \[ v\sim U(-10, 10),\\ \theta\sim N(0,\exp(2v)) \]
## simulate data
set.seed(1234)
NN <- 10000
A <- 10
v <- runif(NN, -A,A)
tau <- exp(v)
theta <- rnorm(NN) * tau
## Gibbs + MH sampling
MH_samples0 <- my_MH_sample0(10000, step_sizes = c(0.5, 0.5))
## JAGS sampling (Gibbs + slice)
library(runjags)
mod.jags <- "
model{
theta ~ dnorm(0, exp(-v))
v ~ dunif(-10,10)
}
"
jags.fit1 <- run.jags(mod.jags, n.chains = 1, sample = 10000, thin = 1,
burnin = 1000, adapt = 500, data = data.frame(x=1),
method = "parallel", modules = 'glm',
monitor=c("theta","v"))
samples <- as.matrix(jags.fit1$mcmc[[1]])
par(mfrow = c(1,3))
plot(theta, v, xlim=c(-20,20), ylim = c(-A,A), pch=20, cex=0.2,main="true")
plot(samples, xlim = c(-20,20),ylim = c(-A,A), pch=20, cex=0.2,main="JAGS")
plot(MH_samples0$theta, MH_samples0$v, xlim = c(-20,20),
ylim = c(-10,10), pch=20, cex=0.2,main="my_MH")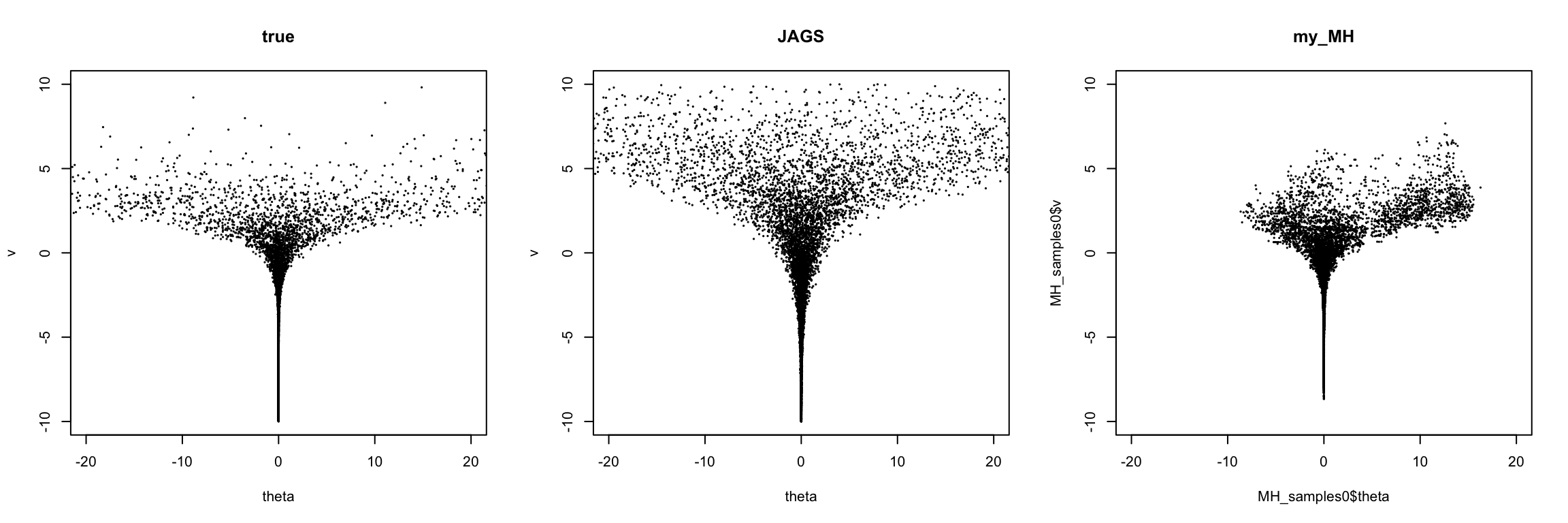 It can be seen that Gibbs + MH performs not well.
It can be seen that Gibbs + MH performs not well.
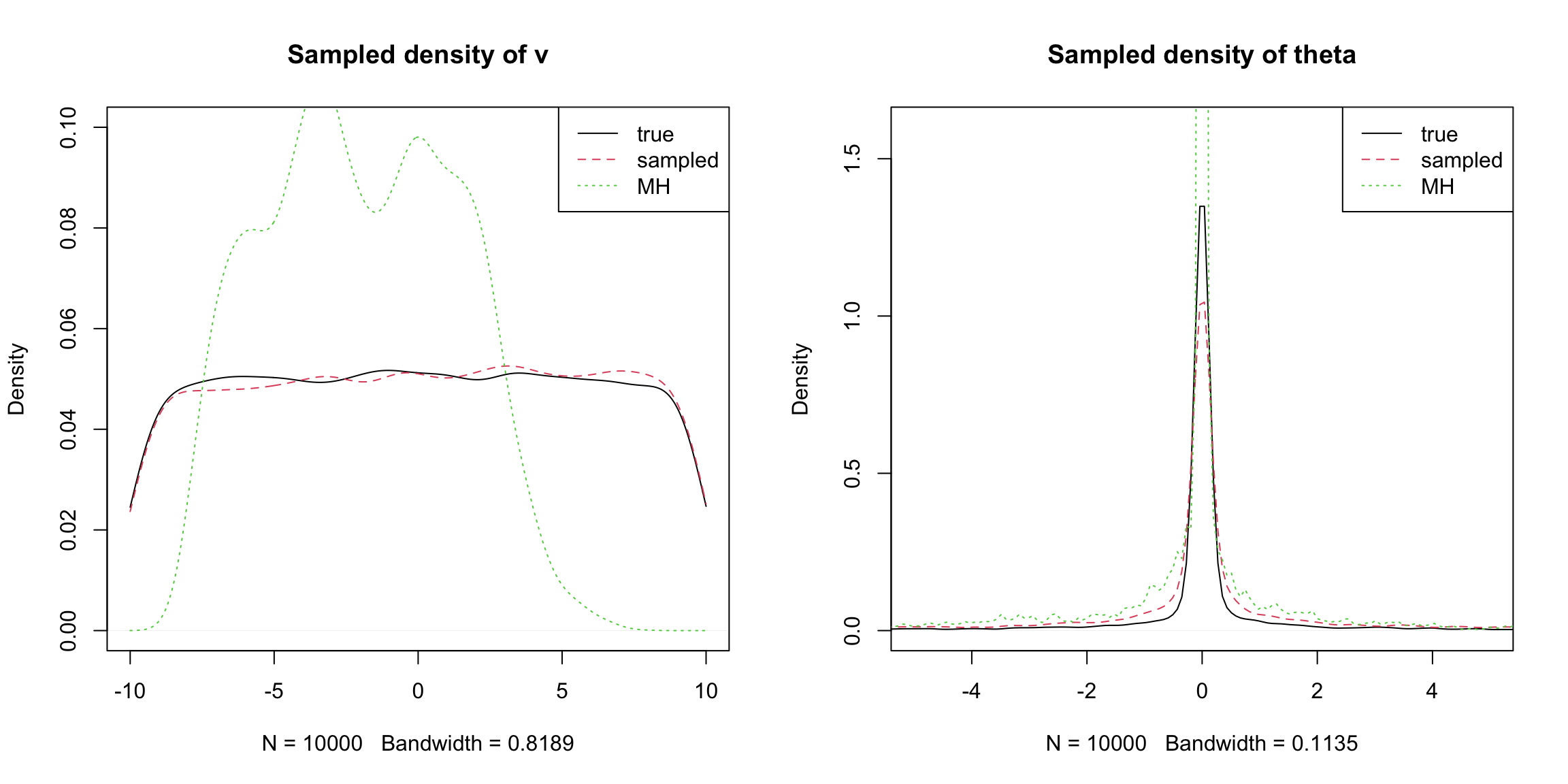
The second example
In the second simulation, we sample the following model: \[ y\sim N(\mu,\sigma^2),\\ \mu\sim N(0,1),\\ \sigma\sim\text{HalfCauchy}(1). \]
library(runjags)
set.seed(1234)
## Sample directly
NN <- 10000
mu = rnorm(NN)
sigma = abs(rcauchy(NN))
y <- mu + rnorm(NN) * sigma
## sample using JAGS
mod.jags <- "
model{
y ~ dnorm(mu, tau)
mu ~ dnorm(0, 1)
sigma ~ dt(0, 1, 1)T(0,)
tau = 1/sigma
}
"
jags.fit1 <- run.jags(mod.jags, n.chains = 1, sample = 10000, thin = 1,
burnin = 1000, adapt = 500, data = data.frame(x=1),
method = "parallel", modules = 'glm',
monitor=c("sigma", "mu", "y"))
samples <- as.matrix(jags.fit1$mcmc[[1]])
## sample using my MH sampler
MH_samples <- my_MH_sampler(10000, step_sizes = c(1, 0.2, 1))
## plot
par(mfrow=c(3,2))
plot(y, mu, xlim = c(-5,5),ylim=c(-4,4),pch=20, cex=0.2, main = "true")
plot(y, sigma, xlim = c(-5,5),ylim=c(0,2.5), pch=20, cex=0.2)
plot(samples[,c(3,2)], xlim = c(-5,5),ylim=c(-4,4), cex=0.2,pch=20, main = "JAGS")
plot(samples[,c(3,1)], xlim = c(-5,5),ylim=c(0,2.5),pch=20, cex=0.2)
plot(MH_samples$y, MH_samples$mu,xlim = c(-5,5),ylim=c(-4,4), pch=20, cex=0.2, main = "my_MH")
plot(MH_samples$y, MH_samples$sigma,xlim = c(-5,5),ylim=c(0,2.5), pch=20, cex=0.2)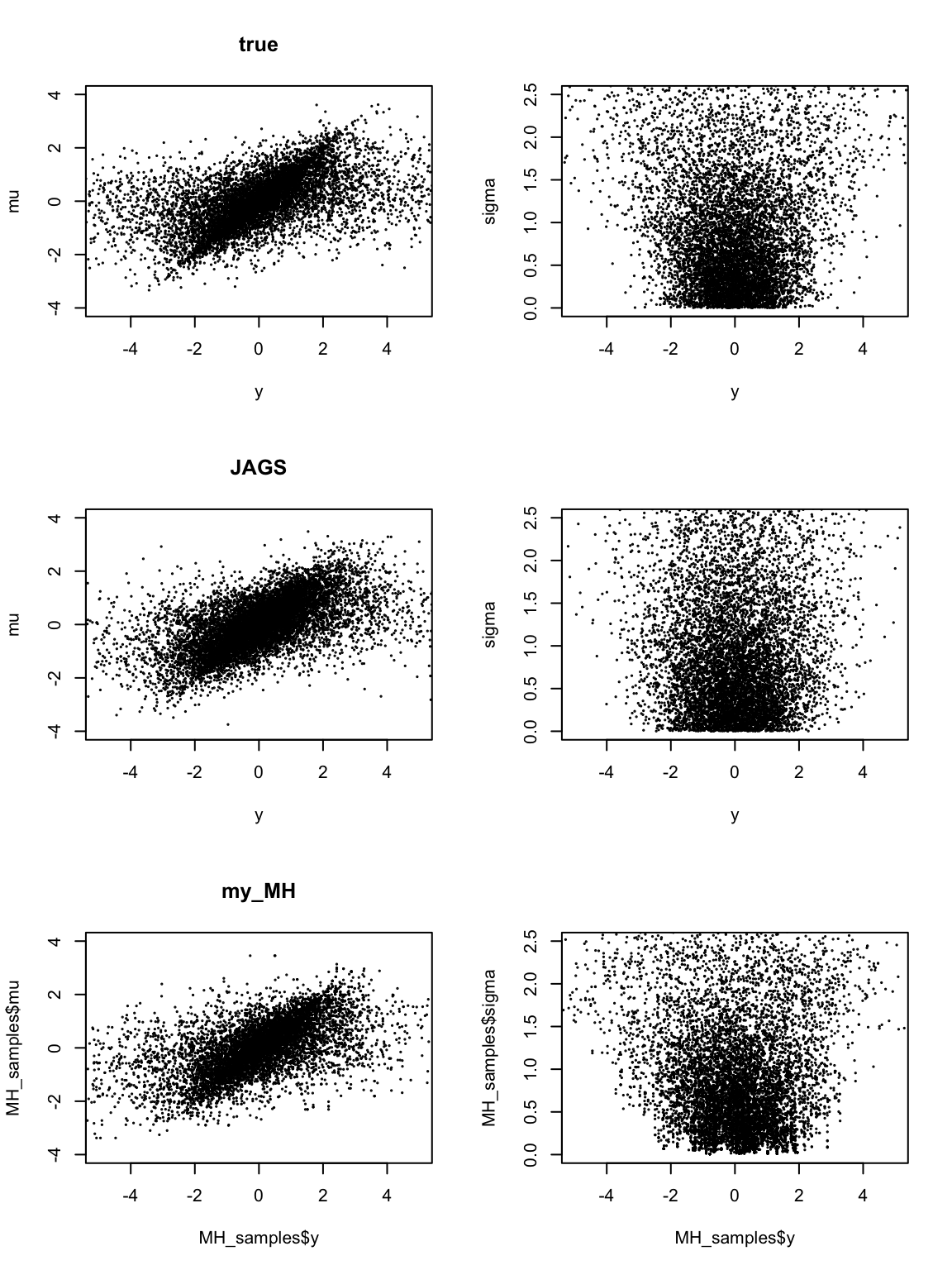
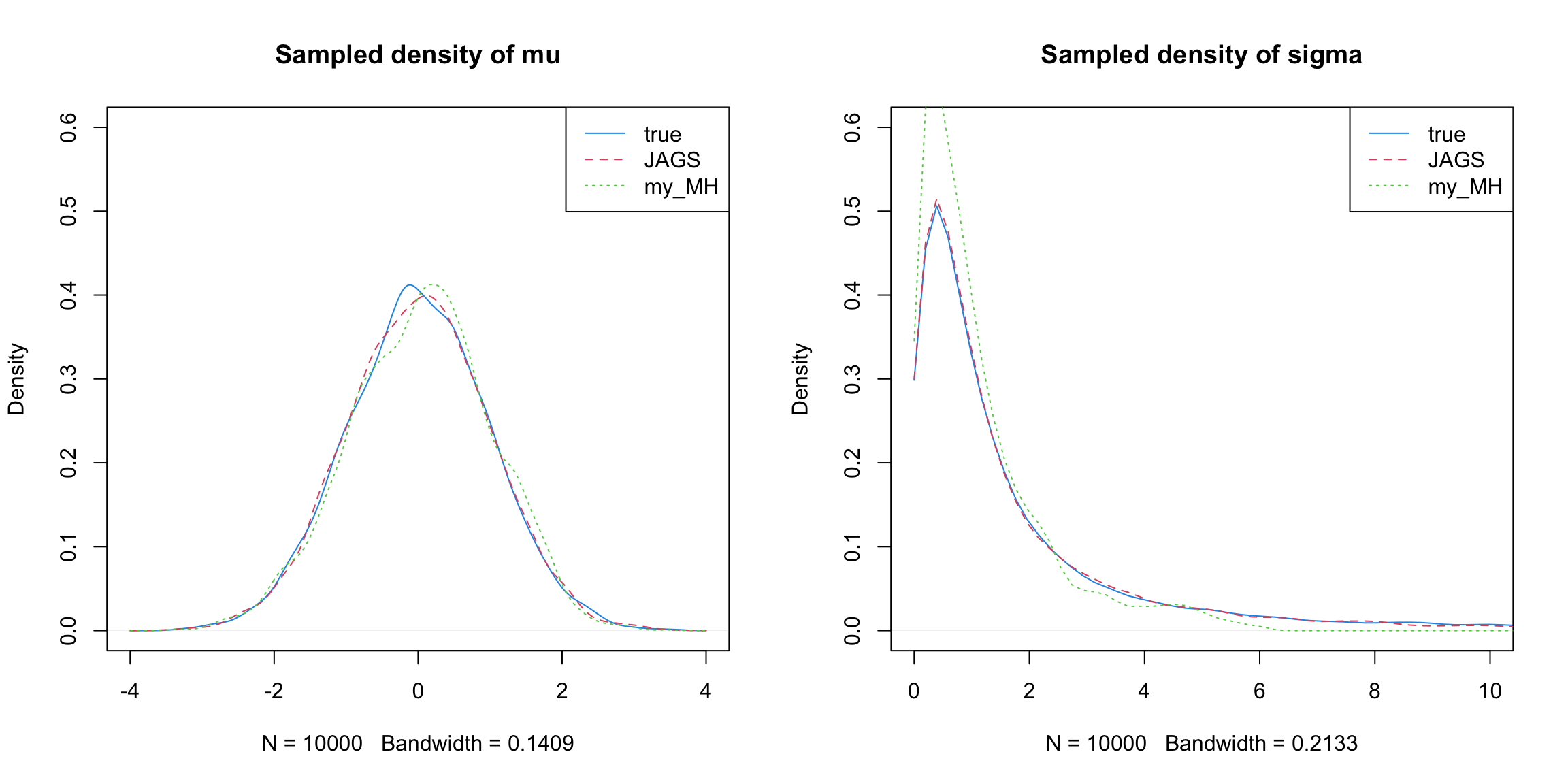
Conclusions
We observe for the above two simple case, only vanilla Gibbs+MH sampler suffers. Gibbs+Slice sampler performs well.
References
- Hierarchical Modeling by Michael Betancourt
- Funneling
- Betancourt and Girolami (2015)